µQFA:
Like QFA, only more awesomer
Conor Lawless
SBSB March 2012
http://lwlss.net/talks/uqfa
What is QFA?
- An extremely high-throughput technology: screens fitness of thousands of independent microbial cultures
- Time-lapse photography of solid agar plates arrayed with up to 308 independent cultures captures rates of change in culture cell density
- Colonyzer gives cell density estimates from pixels
- Dilute culture inoculum allows capture of exponential growth rate from growth curves (fitting logistic model)
- Incubating plates at different temperatures allows investigation of temp. sensitivity (e.g. TS mutations)
- Adding drugs & nutrients to agar or irradiating plates allows investigation of their effects on growth
- Two phenotypes: population growth rate and yield
QFA Growth curves by photographing agar plates

$d(t) = \frac{K d_0 e^{rt}}{K + d_0 \left( e^{rt} - 1\right)}$
Logistic population growth model
QFA epistasis plot - interaction with query mutation

What is µQFA?
- Plain old high-throughput technology for simultaneous screening of the fitness of up to 308 microbial cultures
- Same principles as QFA, but uses microscopy instead of photography to estimate cell density
- Individual cells detected immediately on inoculation
- Growth rate for sick cells which only divide a few times
- Growth rates during both lag and exponential phases
- Growth rate heterogeneity between lineages (cell viability)
- Opportunity for examining fluorescence markers in cell lineages
- Several simultaneous phenotypes: very rich data
Preparing to inoculate a µQFA plate
Loading Cultures
Shaking Cultures
Loading 96 Well Plate
Sterilising Pin Tool
Preparing to inoculate a µQFA plate
Inoculating Cultures Onto Agar
Locating Corner Cultures
Sealing Cultures Into Solid Plate
Automated Stage
Main µQFA experimental components



Main µQFA software components
µQFA timelapse photographs
Microcolony detection (images analysed using OpenCV)
- Single cells at start (with random clumping)
- Rapidly dividing cells out-compete other cells
- Some cells do not divide
Cultures at start of experiment

Cultures at end of experiment

Canny edge detection

Assigning cultures to particles

Assigning cultures to particles

µQFA growth rate distributions

µQFA growth rate distributions

Conclusions
- µQFA experimental protocol & analysis are working
- Tracks cell lineages & provides growth rate estimates immediately after inoculation
- Estimates population growth rate distributions & identifies proportion of viable cells
- Could be used to investigate correlation between growth rate and protein expression
- Validate dynamic, stochastic, population models of growth or statistical models of genetic interaction
Acknowledgements
David Lydall, Peter Banks, Alan Leake 
Raw µQFA images
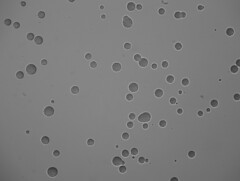



µQFA computer vision
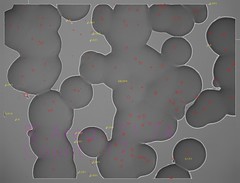

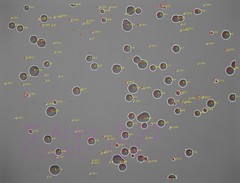

Culture area per pixel in a single QFA photo is much higher than that in µQFA

A common model of microbial growth - evidence for lag phase in µQFA observations?

µQFA "average" growth curves are piecewise linear on log scale


- Detailed modelling of stochastic lag, but with no data:
- Modelling variation in culture volume with observed area:









